A new mechanism for accessing damaged DNA
Advertisement
UV light damages the DNA of skin cells, which can lead to skin cancer. But this process is counteracted by the DNA repair machinery, acting as a molecular sunscreen. It has been unclear, however, how repair proteins work on DNA tightly packed in chromatin, where access to DNA damage is restricted by protein packaging. Using cryo electron microscopy, researchers from the Thomä group at the Friedrich Miescher Institute for Biomedical Research (FMI) have identified a new mechanism whereby repair proteins detect and bind to damaged DNA that is densely packed in nucleosomes.
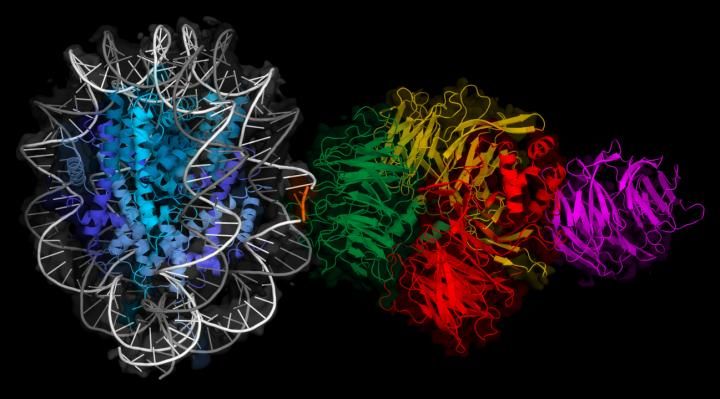
Cryo-EM map of a molecule of UV-DDB (right) binding to DNA wrapped around a histone (left).
FMI
Ultraviolet (UV) light damages DNA, producing small lesions. These UV lesions are first detected by a protein complex known as UV-DDB and - once the lesions have been identified - the rest of the DNA repair machinery swings into action. The question is, how can UV-DDB bind to lesions when the DNA is coiled around the histone protein core of the so-called nucleosome (the basic unit of chromatin - the DNA packaging of eukaryotic chromosomes)?
To gain access, UV-DDB was previously thought to require the assistance of additional proteins that shift the nucleosome. Researchers from the group led by Nicolas Thomä have now found that additional proteins are not necessarily needed to detect UV-induced lesions; instead, the UV-DDB complex takes advantage of the intrinsic dynamics of nucleosomal DNA. The DNA repair factor appears to catch the UV lesions when they are temporarily accessible.
In their study published in Nature, the scientists determined various three dimensional (3D) structures of UV-DDB bound to lesions at different locations around the nucleosome, using cryo-electron microscopy - a technique that allows the 3D structure of biomolecules to be visualized with atomic detail. The researchers concluded that damage detection strategies depend on where the DNA lesion is located. In the case of "accessible" lesions, which can be directly contacted, UV-DDB binds to the lesion tightly. The recognition of "occluded" lesions (facing the histone protein core of the nucleosome) requires additional steps: UV-DDB binds the UV lesions when they are exposed temporarily through the natural dynamics of the nucleosome. One of the lead authors, Syota Matsumoto, explains: "To visualize what happens at the molecular level, imagine a piece of string wrapped around a spool, which becomes accessible when it is pulled forwards or backwards a little bit."
The researchers called the mechanism of DNA damage read-out "slide-assisted site-exposure". This new mechanism operates independently of chromatin remodelers and does not require chemical energy to slide or dislodge nucleosomes.
Thomä comments: "In the past, nucleosomes were thought to be a major obstacle for DNA-binding proteins. In our study, we show that they are not, and that the system is tailored to bind UV lesions wherever they are. What makes this study really powerful is the fact that the mechanism we identified could very well be used by many other types of DNA-binding proteins. Accessing nucleosomal DNA is not only fundamental for DNA repair, but is relevant for all proteins that bind to chromatin. With our study, we define a previously unknown strategy for protein access to chromatinized DNA templates."
Original publication
Syota Matsumoto*, Simone Cavadini*, Richard D. Bunker*, Ralph S. Grand, Alessandro Potenza, Julius Rabl, Junpei Yamamoto, Andreas D. Schenk, Dirk Schübeler, Shigenori Iwai, Kaoru Sugasawa, Hitoshi Kurumizaka, Nicolas H. Thomä; "DNA damage detection in nucleosomes involves DNA register shifting."; Nature (2019).