Bioinformatics platform for the genome-based taxonomical classification of bacteria and archaea
Researchers develop Type Strain Genome Server (TYGS)
Advertisement
"TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy", is the title of the paper published in the current issue of the internationally journal Nature Communications. In this paper, Dr. Jan P. Meier-Kolthoff and Ass. Prof. Dr. Markus Göker, both of the Department for bioinformatics at the Leibniz Institute DSMZ-German Collection of microorganisms and Cell Cultures, share their findings regarding the new Type Strain Genome Server TYGS.

Leibniz-Institut DSMZ
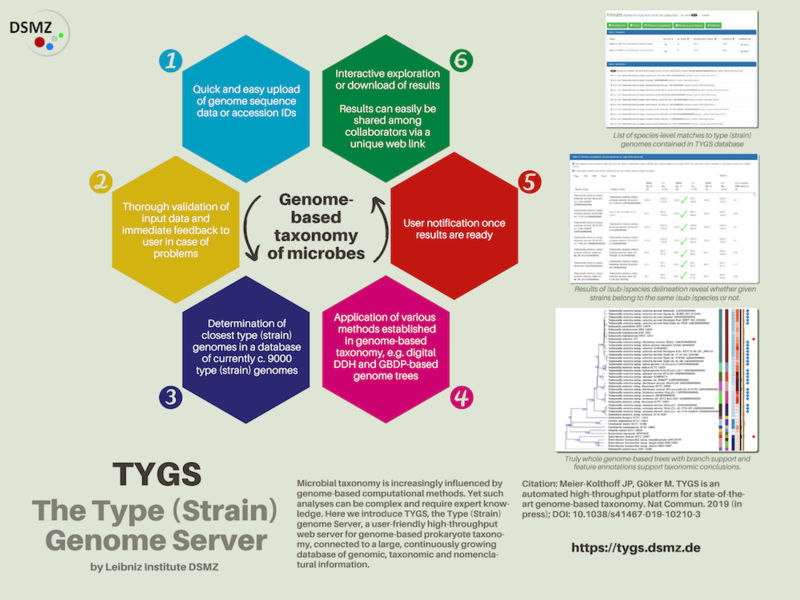
Scientists at the DSMZ developed this Type (Strain) Genome Server as a bioinformatics platform for the genome-based taxonomy of bacteria and archaea. The server is based on a database with weekly updates, which links comprehensive information on taxonomy and nomenclature to almost 9,000 microbial type strain genome sequences and essentially all type strain 16S sequences.
The Type (Strain) Genome Server uses a high-throughput process to automatically determine the type strain genome sequences most closely related to one (or up to several) user-defined genome sequences, and from there calculates, amongst others, a true genome-based family tree, a categorization of species and subspecies using digital DNA:DNA hybridization, and all differences in genomic GC-Content. For this, the TYGS uses a number of scientifically established and frequently cited methods including the Genome-to-Genome Distance Calculator (GGDC) or a phylogenomic process called Genome BLAST Distance Phylogeny (GBDP).
The user receives a specific link to an interactive website of the TYGS-server containing all relevant information in form of tables, modifiable figures ready for publication, references on taxonomy literature as well as complete texts on methods and results. The data can be integrated directly into microbiological studies and papers; in addition, the user is also able to exchange them with cooperation partners via the web link provided. Should the phylogenetic environment be thought to not yet contain genome-sequenced type strains, the user may conveniently use TYGS on the basis of this result to request a more comprehensive tree analysis of the 16S rRNA gene sequences.