On the trail of cancer stem cells
What goes on inside and between individual cells during the very earliest stages of tumor development?
Advertisement
single cell sequencing technologies and a mouse model have enabled researchers to comprehensively map the cellular diversity of whole salivary gland tumors and trace the path of cancer stem cells.
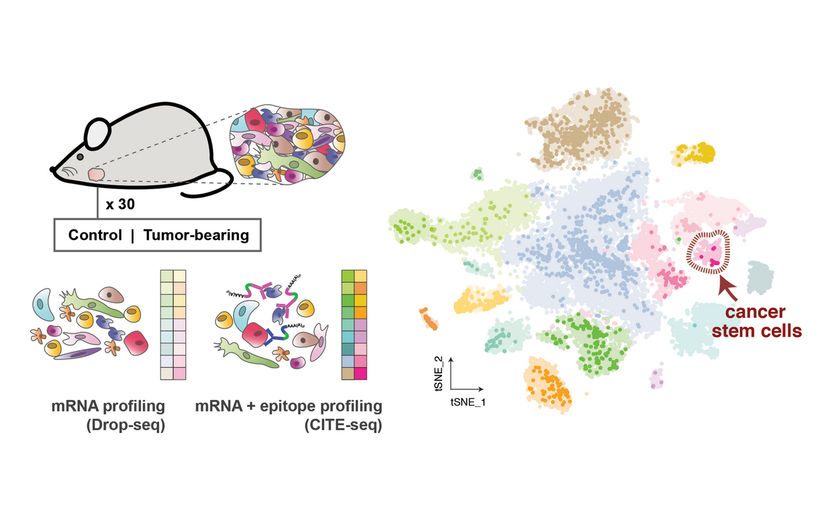
Left: The researchers individually sequenced more than 26,000 cells from healthy mice and mice with salivary gland tumors. They used different strategies to analyze both the RNA inside the cells (mRNA profiling) and the proteins on the surface of cells (epitope profiling) to identify the cells. Right: The result was a comprehensive cell atlas of the healthy and tumor-bearing salivary gland (shown here) that revealed distinct populations of the different cell types, including a very small population of cancer stem cells.
© Samantha Praktiknjo, MDC
Two research teams from the Max Delbrück Center for Molecular Medicine and their collaborators have produced a detailed cell atlas of an entire salivary gland tumor in a mouse model, mapping individual cells throughout the tumor and its surrounding tissue. The “single cell” approach, recently described in Nature Communications, has provided key insights about cellular composition changes through the earliest stages of cancer development.
A solid tumor is not, as many might assume, a lump of cells that are all the same. Rather it is mix of many different cell types, including a variety of stromal and immune cells besides the actual tumor cells.
“Conventional methods in molecular biology often consider a sample as a whole, which fails to recognize the complexity within it,” said Dr. Samantha Praktiknjo, senior scientist and first author from MDC’s Systems Biology of Gene Regulatory Elements Lab headed by Professor Nikolaus Rajewsky at the Berlin Institute for Medical Systems Biology (BIMSB). Developing a detailed understanding of the different cells within a tumor and how they interact could help identify more effective treatment strategies.
Strength in numbers
The team used single-cell RNA sequencing technologies developed in the Rajewsky lab and novel epitope profiling to produce the cell atlas, and identified the cells that were specific to the tumor by leveraging the reproducibility and the large sample size of their data.
The latter was possible by using a mouse model, developed in MDC’s Signal Transduction in Development and Cancer Lab headed by Professor Walter Birchmeier, which harbors designed mutations that induce a salivary gland squamous cell carcinoma. This system provides a consistent supply of genetically similar tumors to sequence from the earliest stages of development, which is nearly impossible with human patients.
“In a patient, the tumor is already developed and you cannot go back and rewind time and look at how it started,” said Dr. Benedikt Obermayer, a co-first author now at the Berlin Institute of Health (BIH ). “Here, we have a model that is so controlled, we can watch it happen.” And Dr. Qionghua Zhu, the third first author and a former postdoc at the Birchmeier Lab, added: “To fight cancer effectively, we need to find the driver mutations. This method gives us clues about the evolution trajectories of a tumor.”
Sequencing technologies have advanced so that it is now possible to quickly and affordably sequence the RNA inside single cells, one at a time, as well as the proteins on the surfaces of cells in the tissues. While other methods grind up the tissue and identify what genes and molecules are present in the mix, the single cell approach precisely identifies how many of each type of cell is present, and which genes and molecules are associated with which cell.
For this study, the researchers sequenced more than 26,000 individual salivary gland cells from mice with tumors and healthy mice. They used computational models to analyze the huge amount of data and identify each individual cell and sort them into groups – such as stromal cells, immune cells, saliva producing cells, cancer cells – based on the hundreds of genes expressed and molecules present.
A surprise
The single cell approach revealed something that surprised the researchers: “When I saw the data, I thought, where is the tumor?” Obermayer said. The population of cancer stem cells in the tumor was extremely small – less than one percent of all profiled cells in the tissue. Due to their low abundance, investigation of these cells still heavily depends on assumptions about surface markers and is often performed in cell culture-based systems. Here, the authors were able to identify the cancer stem cells directly from the solid tumor samples with their single cell approach.
Furthermore, the team was able to predict the progression of the different cell types through different stages of tumor development. Their model suggests that the cancer stem cells emerge from cancerous basal cells, then develop into another subtype before ultimately becoming a population of cells similar to luminal cells, a cell type present in normal, healthy salivary glands.
This progression supports the idea that when something goes awry in the basal cells of this solid tumor model, they are triggered to turn into cancer stem cells, which can then become a different type of cell. “What I found fascinating was clearly seeing the order of signals and events, transitioning from the progenitor to the progeny populations of the cancer stem cells,” Praktiknjo said.
Next steps
Further research is required to verify that individual cells are transforming through these stages, and explore the cellular and molecular interactions driving tumor growth. The team anticipates the approach they’ve demonstrated here can be applied to other cancer types as well.
“To me the main conceptual insight is that we can apply ideas from single-cell based developmental biology to reconstruct molecular progression of tumorigenesis ,” said Professor Nikolaus Rajewsky, who heads MDC’s Systems Biology of Gene Regulatory Elements Lab and is the scientific director of the BIMSB.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Cell Analysis
Cell analyse advanced method allows us to explore and understand cells in their many facets. From single cell analysis to flow cytometry and imaging technology, cell analysis provides us with valuable insights into the structure, function and interaction of cells. Whether in medicine, biological research or pharmacology, cell analysis is revolutionizing our understanding of disease, development and treatment options.
Topic World Cell Analysis
Cell analyse advanced method allows us to explore and understand cells in their many facets. From single cell analysis to flow cytometry and imaging technology, cell analysis provides us with valuable insights into the structure, function and interaction of cells. Whether in medicine, biological research or pharmacology, cell analysis is revolutionizing our understanding of disease, development and treatment options.