OSCAR detects cells in standby mode
New tool to better understand the biology of dormant stem cells
Advertisement
dormancy is a sleep-like state of cells that protects them from genetic damage and thus prolongs their survival. This state is reversible and characterized by low metabolic activity and division rate. Researchers from the Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) and the University of Oxford, UK, developed a method to detect dormancy in real time. Using the fluorescence measurement OSCAR (Optical Stem Cell Activity Reporter), cell populations with different dormancy values could be identified and isolated in the small intestine model. OSCAR is thus a suitable tool to better understand the biology of dormant stem cells.
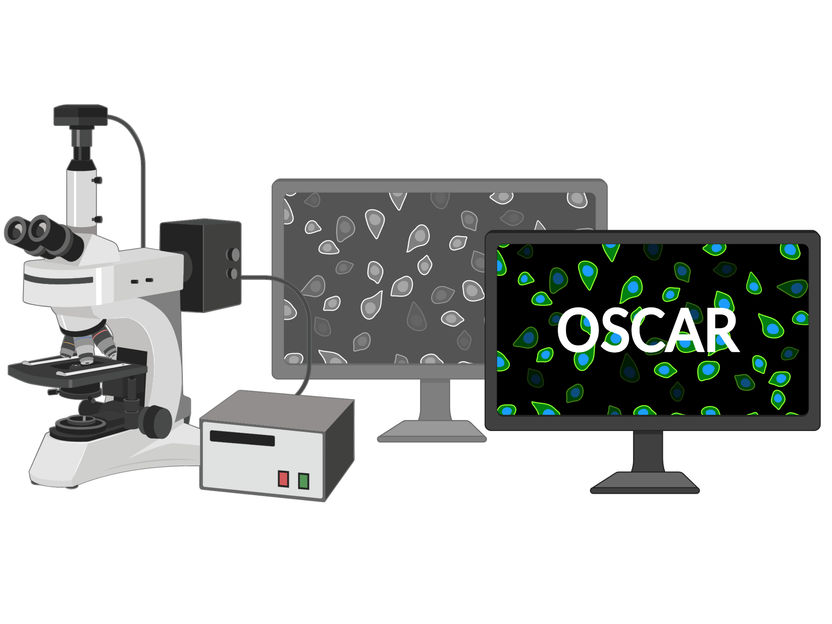
OSCAR (optical stem cell activity reporter) can be used to detect dormant stem cells in tissue. A short peptide is incorporated into the backbone of a fluorescent protein. Due to the lack of phosphorylation, an increase in green fluorescence is observed.
FLI / Kerstin Wagner & Francesco Neri, created with BioRender.com
An important example of dormant cells are somatic stem cells, which are characterized by their potential for differentiation and self-renewal. They have a high transplantation potential, are more resistant to cellular stress and have a great potential to be used in regenerative medicine. Hematopoietic stem cells in bone marrow, for example, are needed only when injured and otherwise exist in a dormant state. Similarly, in cancer, relapse after an apparently successful therapy, is thought to be caused by activation of long-term dormant cancer stem cells.
This cellular resting state is called dormancy. It is reversible and characterized by reduced metabolic activity, cell division and mRNA synthesis. This sleep-like state (standby mode) thus protects the cell from genetic damage, prolongs its survival, and is critical for tissue homeostasis and cellular response to injury or transplantation. Dormant cells are found in numerous tissues and organs. However, their identification, isolation and characterization, independent of the tissue of origin, is difficult. Researchers at the Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) in Jena, Germany, in cooperation with partners from the University of Oxford, UK, have now developed a fluorescence measurement method to detect and isolate dormant cells in specific cell types in real time.
Difficulties in the detection of resting cells
"Somatic stem cells are usually identified by surface markers, fluorescent proteins or via labelable cells. However, these methods only label specific stem cell lines are only valid for a certain species type and are technically very challenging," explains Dr. Rasmus Freter from FLI, first author of the study recently published in Nature Communications. "A reliable, specific and, above all, conserved marker for quiescent stem cells that allows their visualization and isolation has not yet been found." Moreover, isolating these cells using surface markers requires prior knowledge, which precludes characterization of previously unknown stem cell populations from the outset. The Jena researchers therefore sought a method to more easily detect dormant cells.
Low RNApII-Ser2 phosphorylation is characteristic of dormant stem cells
"It has been known for more than 10 years that dormant stem cells have a lower amount of total RNA," explains Dr. Francesco Neri, research group leader at FLI and now Associate Professor at the University of Turin. mRNA is transcribed by RNA polymerase II (RNApII), which is phosphorylated to initiate mRNA transcription. High level of RNApII phosphorylation are detected in proliferating and differentiated active cells, while the low RNApII-Ser2 phosphorylation is highly specific to quiescent cells and distinguishes them from other cell types. "We examined various tissues for adult stem cells that had low levels of RNApII-Ser2 phosphorylation," Prof. Neri reports, "and were thus able to detect cells without detectable RNApII-Ser2 phosphorylation in all tissues examined, including brain, hematopoietic stem cells, and small intestine."
Development of a fluorescent kinase reporter - OSCAR
"In a further step, we exploited this observation and specifically looked for a fluorescent reporter that could distinguish between cells with high or low RNApII-pSer2 kinase activity, in order to be able to develop a tool for identifying and isolating living resting cells from any tissue," said Dr. Rasmus Freter, explaining the next steps.
The researchers achieved this with a ratiometric fluorescent optical stem cell activity reporter, called OSCAR, based on the fact that phosphorylation of RNApII is largely absent in quiescent stem cells of different lineages. "Proper visualization of kinase activity in living cells has long been a major challenge," Dr. Freter points out. "For example, most kinase reporters use FRET technology, but this is technically challenging and highly susceptible to high background signals. In addition, the dynamic ranges for signal-to-noise ratios are about 20-30%, which is much too low for isolating cells by fluorescence-activated cell sorting, FACS."
To overcome these technical barriers and develop a sensitive yet practical tool, Prof. Neri's researchers designed a genetically encoded kinase reporter of the relevant RNApII Ser2 kinase. They incorporated a short peptide containing a specific phosphorylation acceptor site into the backbone of a fluorescent protein. "In this context, unphosphorylated peptide insertion did not affect fluorescence performance, while phosphorylation of the inserted peptide reduced fluorescence. As a result, high fluorescence would positively indicate low kinase activity and thus mark RNApII-pSer2-low resting cells," Prof. Neri said. Mass spectrometry analysis confirmed the specificity of the phosphorylation.
The kinase reporter, referred to as "OSCAR," has a significantly improved dynamic range compared with previous FRET-based systems and thus can also be used for FACS sorting of dormant intestinal cells and time-lapse microscopy. For example, using the small intestinal crypt as a model, OSCAR allowed real-time detection of the dynamics of dormancy induction and cellular differentiation in vitro. Furthermore, OSCAR succeeded in identifying and isolating multiple populations of different OSCAR states - from high to low - in the living organism (in vivo).
So far, no toxic effects of OSCAR expression have been detected in mammalian cells and mice. "We therefore anticipate that OSCAR could prove to be a useful tool in characterizing quiescent cells both in vitro and in vivo," said Prof. Neri, summarizing the study results. OSCAR is thus not only a suitable tool to better understand the biology of dormant stem cells. It is also possible that this new method will make it easier to detect suspected dormant cancer stem cells in a living cell population and thus remove them at an early stage.