Hurdle of microscopy overcome
Smallest cell structures can now be imaged even better
Advertisement
High-resolution microscopy makes it theoretically possible to image cell structures with a resolution of a few nanometres. However, this has not yet been possible in practice.
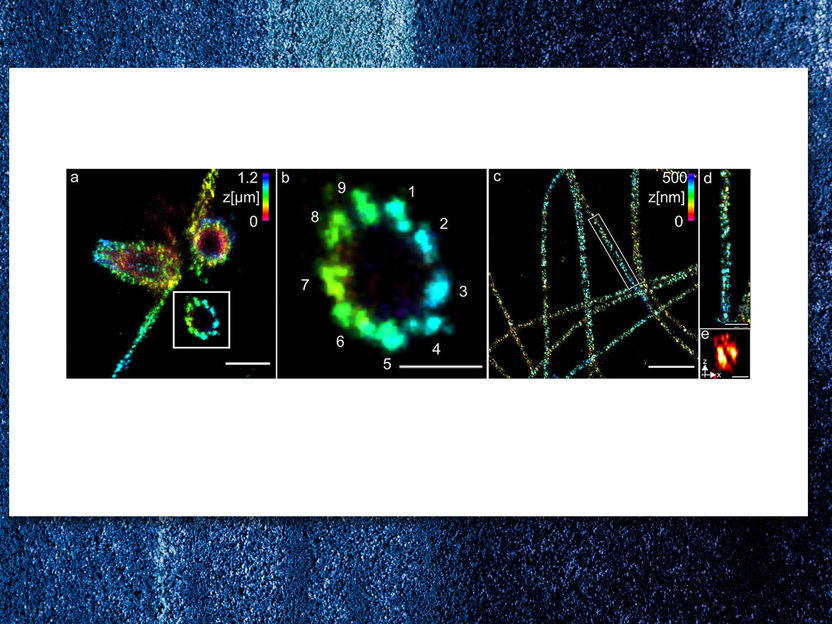
(a) Three-dimensional Ex-dSTORM of 3.2 times expanded centrioles. Measuring bar one micrometer. (b) The enlarged section of (a) shows the nine-fold symmetry of the percentile. Measuring bar 500 nanometers. (c) Three-dimensional Ex-dSTORM of 3.1-fold expanded tubulin filaments. Measuring bar two microns. (d) The magnification in (c) shows a tubulin filament; measuring bar 500 nanometers. (e) The cross-section of a tubulin filament shows its hollow structure. Measuring bar 200 nanometres.
Team Markus Sauer / Universität Würzburg
The reason for this is that antibodies carrying a fluorescent dye are usually used to mark the cell structures. The dye is therefore not located directly at the target structure, but about 17.5 nanometers away. Partly because of this distance error, the theoretically achievable resolution could not be achieved so far.
Publication in Nature Communications
An international research team has now overcome this hurdle. This was achieved by combining the high-resolution fluorescence microscopy dSTORM with the expansion microscopy ExM. The journal Nature Communications presents the results.
The publication was led by a team from the Biocenter of the Julius Maximilian University (JMU) Würzburg: Professor Markus Sauer, Head of the Department of Biotechnology and Biophysics, with PhD students Fabian Zwettler and Sebastian Reinhard. Professors Paul Guichard from the University of Geneva (Switzerland) and Toby Bell from Monash University (Australia) also played a key role.
Obstacles to combining dSTORM and ExM
The dSTORM method, which was developed in Professor Sauer's group, achieves an almost molecular resolution of about 20 nanometers. In order to further increase the resolution, a combination with expansion microscopy, which has been available for a few years, seemed promising.
In ExM, the sample to be examined is crosslinked into a swellable polymer. The interactions of the molecules in the sample are then destroyed and the sample is allowed to swell with water. This leads to an expansion: the molecules to be imaged drift spatially apart by a factor of four.
Why the two methods could not be combined so far:
-
The fluorescent dyes used by dSTORM to label the molecules did not survive the polymerization of the aqueous gel.
For dSTORM, a buffer solution is necessary, but the expanded sample could be contracted again.
Distance error significantly reduced
"By stabilizing the gel and immune staining only after expansion, we could overcome these hurdles and successfully combine the two microscopy methods," says Markus Sauer happily. As a result, the distance error melts to just five nanometers when expanded 3.2 times. This makes fluorescence imaging with molecular resolution possible for the first time.
The researchers have shown how well their method works with centriols and structures that are composed of the protein tubulin. Among other things, they were able to visualise tubulin tubes as hollow cylinders with a diameter of 25 nanometres. The researchers succeeded in sharply imaging groups of three made up of tubulin structures at a distance of 15 to 20 nanometres at the centrioles. The centrioles are cell structures that play an important role in cell division.
Professor Sauer concludes: "For many important cell components, the combination of ExM and dSTORM now allows us to gain detailed insights into molecular function and architecture for the first time. The team therefore plans to apply the method to different structures, organelles and multiprotein complexes of the cell.
Note: This article has been translated using a computer system without human intervention. LUMITOS offers these automatic translations to present a wider range of current news. Since this article has been translated with automatic translation, it is possible that it contains errors in vocabulary, syntax or grammar. The original article in German can be found here.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic world Fluorescence microscopy
Fluorescence microscopy has revolutionized life sciences, biotechnology and pharmaceuticals. With its ability to visualize specific molecules and structures in cells and tissues through fluorescent markers, it offers unique insights at the molecular and cellular level. With its high sensitivity and resolution, fluorescence microscopy facilitates the understanding of complex biological processes and drives innovation in therapy and diagnostics.
Topic world Fluorescence microscopy
Fluorescence microscopy has revolutionized life sciences, biotechnology and pharmaceuticals. With its ability to visualize specific molecules and structures in cells and tissues through fluorescent markers, it offers unique insights at the molecular and cellular level. With its high sensitivity and resolution, fluorescence microscopy facilitates the understanding of complex biological processes and drives innovation in therapy and diagnostics.