A bucket full of genes: pond water reveals tropical frogs
Advertisement
When a frog jumps into a pond, it inevitably leaves behind traces of its genetic material. Using water samples from the Bolivian lowlands, Senckenberg scientists were now able to demonstrate that the analysis of this so-called environmental DNA allows the reliable identification of the frog species that inhabit a body of water. In their feasibility study, the researchers show that the analysis of environmental DNA can serve as a cost-effective alternative to traditional survey methods in species-rich regions and may speed up the necessary world-wide inventory of biological diversity.
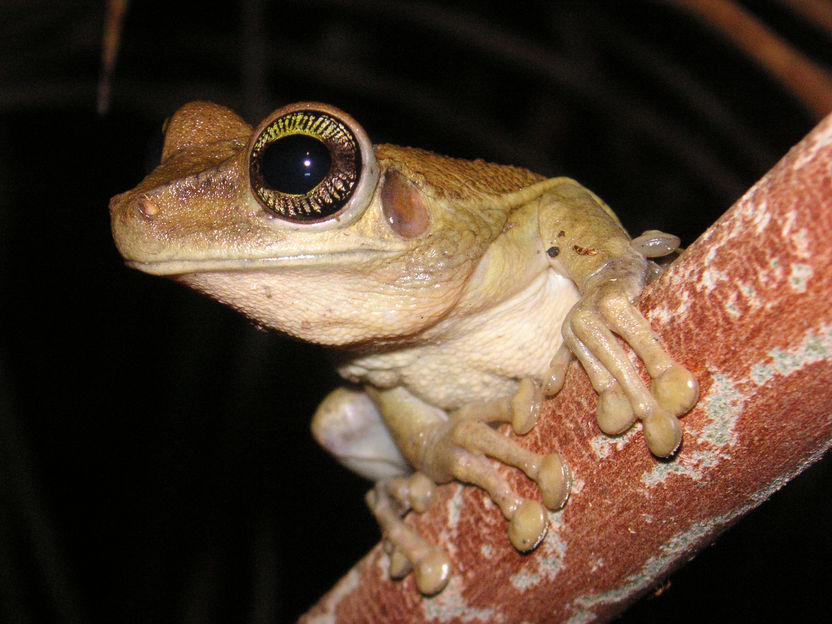

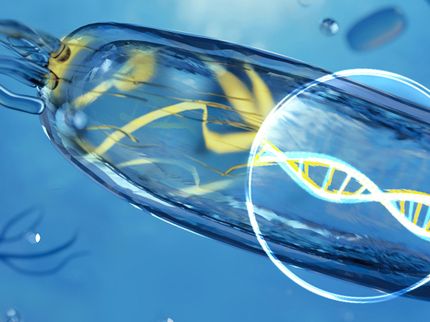
The presence of the frog species Osteocephalus taurinus was only detectable using eDNA.
Copyright: Martin Jansen
Globally, there are almost 7,000 species of frogs, the majority of which occur in the tropics. In order to systematically survey their distribution and detect population trends, experts until recently had to stake out the amphibians – a time-consuming and costly task. Senckenberg scientists now show that there may be a simpler way.
Their method is based on the fact that all living beings leave behind traces of DNA, the so-called environmental DNA or eDNA. “When a frog jumps into a pond, it sheds minute skin particles or other tissue. A water sample will therefore contain a collection of organic material from the frogs that inhabited the pond. The frogs’ genetic material can be isolated from this medley and compared to the database in order to document the species that are present”, explains Dr. Miklós Bálint of the Senckenberg Biodiversity and Climate Research Centre.
As shown by Bálint and his colleagues in a recent feasibility study, two liters of pond water are sufficient to apply this method. The team collected water samples from five ponds in the Bolivian savanna and subsequently isolated and sequenced the frog DNA contained in the samples. In the process, the researchers discovered genetic traces that could be assigned to 25 species of frogs.
“In parallel with this method, we also identified frogs in the traditional manner through observation and analysis of their calls. A comparison shows that both methods have a very similar success rate”, according to Dr. Martin Jansen, a herpetologist at the Senckenberg Research Institute and Nature Museum in Frankfurt.
In fact, six frog species were only identified due to the genetic material they left behind. Bálint comments as follows: “Based on the eDNA in the pond water, we were able to prove the presence of frog species that we failed to observe or identify based on their calls – for example, because they were still in their larval stage or since it only involved single individuals. In those cases, eDNA can provide a more exact picture.”
The team used a detailed cost analysis to document that for many study areas in species-rich tropical regions it would be more cost-effective to inventory species diversity by means of eDNA. Despite the high initial costs of the DNA analysis, the effort required to properly train experts and then dispatch them to often very remote regions to conduct observations is disproportionally higher.
According to the study’s authors, eDNA therefore constitutes a significant step toward a global inventory of biological diversity. “To date, there are very few comprehensive scientific surveys regarding the occurrence of organisms. I envision that we could take water samples from 10,000 ponds in the rainforest and the savanna and use environmental DNA to study the occurence of frogs at a hitherto unprecedented level of detail”, explains Bálint.
This survey is of high importance, since – along with many other species of plants and animals – frogs face a particularly high risk of extinction due to the global change. More than one-third of all species are already considered threatened, and the trend is rising. “However, we will only be able to protect these species if we know where they occur. Our taxonomic expertise aids us in recognizing new species, while the analysis of eDNA renders it more efficient to inventory already known species in many locations. Both methods thus optimally complement each other”, adds Jansen in closing.
Original publication
Bálint, M. et al.; "Accuracy, limitations and cost-efficiency of eDNA-based community survey in tropical frogs"; Molecular Ecology Resources; 2018