Controlled Nano-Assembly
Universal pH regulated assembly of DNA nanostructures
Advertisement
DNA, the carrier of genetic information, has become established as a highly useful building material in nanotechnology. One requirement in many applications is the controlled, switchable assembly of nanostructures. Scientists have now introduced a new strategy for control through altering ph value. It is based on ethylenediamine, which only supports the assembly of DNA components in a neutral to acidic environment—independent of the base sequences and without metal ions.
© Wiley-VCH
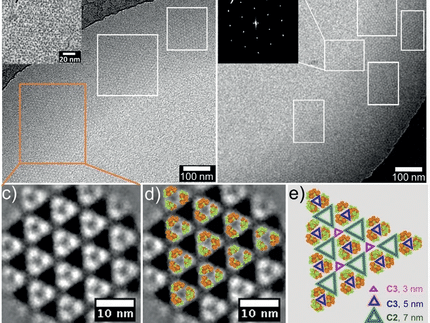
A set of short single strands of DNA can be knitted into a tile which further aggregates into a rich range of geometries through self-directed, sticky-end cohesions. The so-called tile assembly mimics a crystal-forming process in nature. Once the right strands are designed, a specific structure forms through a self-assembly process. Researchers hope to use this method in the future to organize nanomaterials or make nanorobots that could carry out tiny interventions in diseased organs or even individual cells. Nanoelectronics and nanocatalysis are also possible areas of future application.
The controlled, switchable assembly of nanostructures occurs with the help of certain DNA structural motifs that change their shape when the pH changes. However, these structures are based on very specific base sequences. In contrast, a sequence-independent method would offer access to a universal, versatile method for the self-assembly of DNA, significantly broadening the possible areas of application for dynamic DNA nanotechnology.
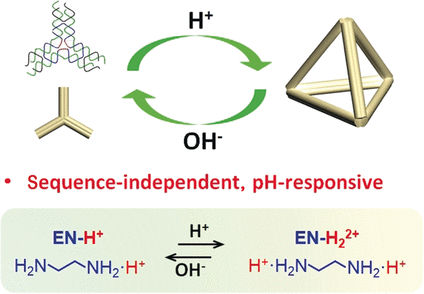
Scientists at Hefei University of Technology; University of Science and Technology of China, Hefei, China, and Purdue University in West Lafayette, USA, start off with a small organic molecule, ethylenediamine, H2N—CH2—CH2—NH2. In water, one or both of the amino groups (–NH2) reversibly bind an additional proton (H+), depending on the pH value. The relative amounts of the three possible species depend strongly on the pH. The ethylenediamine molecules with a twofold positive charge are able to electrostatically shield the negative charges in the DNA backbone so they repel each other less, which favors self-assembly.
The team headed by Yulin Li, Zhaoxiang Deng, and Chengde Mao was able to use an ethylenediamine-containing buffer to assemble individual cross-shaped DNA building blocks (tiles) into an extended two-dimensional honeycomb structure in a neutral to acidic environment. In slightly alkaline conditions, no extended structures formed. As an example of a three-dimensional structure, the researchers made tetrahedral cages out of building blocks shaped like three-pointed stars. At a pH of 6.5, the stars were reversibly converted to tetrahedron; at a pH of 8 they reverted to the star shape or individual strands.
To shield the negative charges in the DNA, positively charged metal ions like Mg2+ are usually used. The ethylenediamine buffer requires no metal ions. Metal ions increase the activity of many enzymes, particularly DNA-splitting nucleases. This new system is thus better suited for use in the presence of enzymes.
Original publication
Yongfei Li, Lei Song, Bang Wang, Prof. Jianbo He, Prof. Yulin Li, Prof. Zhaoxiang Deng, Prof. Chengde Mao; "Universal pH‐Responsive and Metal‐Ion‐Free Self‐Assembly of DNA Nanostructures"; Angew. Chem.; 2018
Yongfei Li, Lei Song, Bang Wang, Prof. Jianbo He, Prof. Yulin Li, Prof. Zhaoxiang Deng, Prof. Chengde Mao; "Universal pH‐Responsive and Metal‐Ion‐Free Self‐Assembly of DNA Nanostructures"; Angew. Chem. Int. Ed.; 2018