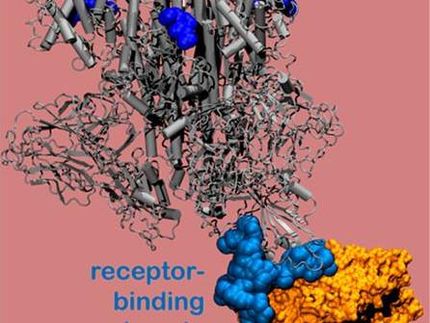
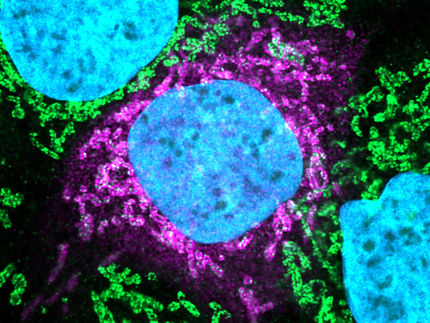
Likely Entrypoints for SARS-CoV-2
Two specific cell types in the nose have been identified as likely initial infection points for the novel coronavirus
Advertisement
Using data from the Human Cell Atlas, scientists discovered that goblet and ciliated cells in the nose have high levels of the entry proteins that SARS-CoV-2 uses to get into our cells.
Tumisu, pixabay.com, CC0
The identification of these cells by researchers from the Wellcome Sanger Institute, University Medical Centre Groningen, University Cote d’Azur and CNRS, Nice and their collaborators, as part of the Human Cell Atlas Lung Biological Network, could help explain the high transmission rate of SARS-CoV-2.
Reported in Nature Medicine, this first publication with the Lung Biological Network is part of an ongoing international effort to use Human Cell Atlas data to understand infection and disease. It further shows that cells in the eye and some other organs such as the heart also contain the viral-entry proteins. The study also predicts how a key entry protein is regulated with other immune system genes and reveals potential targets for the development of treatments to reduce transmission.
Novel coronavirus disease - COVID-19 – primarily affects the lungs and airways. Patient’s symptoms can be flu-like, including fever, coughing and sore throat, while some people may not experience symptoms but still have transmissible virus. In the worst cases, the virus causes pneumonia that can ultimately lead to death. The virus is thought to be spread through respiratory droplets produced when an infected person coughs or sneezes, and appears to be easily transmitted within affected areas. So far the virus has cost more than 176,000 lives.
Pinpointing the cell types involved in the infection
Scientists around the world are trying to understand exactly how the virus works, to help prevent transmission and develop a vaccine. While it is known that the virus that causes COVID-19 disease, known as SARS-CoV-2, uses a similar mechanism to infect our cells as a related coronavirus that caused the 2003 SARS epidemic, the exact cell types involved in the nose had not previously been pinpointed.
To discover which cells could be involved in COVID-19, researchers analysed multiple Human Cell Atlas (HCA) consortium datasets of single cell RNA sequencing, from more than 20 different tissues of non-infected people. These included cells from the lung, nasal cavity, eye, gut, heart, kidney and liver. The researchers looked for which individual cells expressed both of two key entry proteins that are used by the virus to infect our cells.
Dr Waradon Sungnak, the first author on the paper from Wellcome Sanger Institute, said: “We found that the receptor protein - ACE2 - and the TMPRSS2 protease that can activate SARS-CoV-2 entry are expressed in cells in different organs, including the cells on the inner lining of the nose. We then revealed that mucus-producing goblet cells and ciliated cells in the nose had the highest levels of both these virus proteins, of all cells in the airways. This makes these cells the most likely initial infection route for the virus.”
Cells in the nose are highly accessible to the virus
Dr Martijn Nawijn, from the University Medical Center Groningen in the Netherlands, said, on behalf of the HCA Lung Biological Network: “This is the first time these particular cells in the nose have been associated with COVID-19. While there are many factors that contribute to virus transmissibility, our findings are consistent with the rapid infection rates of the virus seen so far. The location of these cells on the surface of the inside of the nose make them highly accessible to the virus, and also may assist with transmission to other people.”
The two key entry proteins ACE2 and TMPRSS2 were also found in cells in the cornea of the eye and in the lining of the intestine. This suggests another possible route of infection via the eye and tear glands, and also revealed a potential for fecal-oral transmission.
When cells are damaged or fighting an infection, various immune genes are activated. The study showed that ACE2 receptor production in the nose cells is probably switched on at the same time as these other immune genes.
ACE2 can also be found in the heart
Up to 20 per cent of hospitalized COVID-19 patients also suffer myocardial damage and consequent cardiac decompensation. Thus, it was crucial to map SARS-CoV-2 receptor and enabling proteases gene expression in the heart as well. “We analysed ~500,000 single cells from 14 human hearts and identified three cellular compartments expressing the entry receptor: pericytes, cells of the heart fine capillary network; cardiac muscle cells; and fibroblasts, the cells contributing to maintain the heart structure”, said Dr Michela Noseda from the National Heart & Lung Institute at Imperial College, London. “Knowing the exact target cells of the virus in the heart provides the basis to understand the mechanisms of damage and guide treatment choices.”
“We have an exceptional set of single cell data”, said Professor Norbert Hübner, head of the Genetics and Genomics of Cardiovascular Diseases group at the Max Delbrück Center for Molecular Medicine (MDC) who also heads projects at the Berlin Institute of Health (BIH ) and German Center for Cardiovascular Diseases (DZHK ). Together with Dr Jonathan Seidman, the Bugher Professor of Cardiovascular Genetics at Harvard Medical School, he coordinates a team of 13 scientists from Germany, the United Kingdom, and the United States dedicated to probing and understanding the human heart, cell by cell. Michela Noseda and Sarah Teichmann are part of this team. “We found the ACE2 receptor in particular in the pericytes. This receptor probably has a fundamental role in maintaining blood flow in the body. However, its role in cardiac problems of COVID-19 patients is another matter. We still don’t know whether the cardiac damage is caused by the virus itself or whether it is a secondary effect.”
Using the Human Cell Atlas to understand COVID-19
The work was carried out as part of the global Human Cell Atlas consortium which aims to create reference maps of all human cells to understand health and disease. More than 1,600 people across 70 countries are involved in the HCA community, and the data is openly available to scientists worldwide.
Dr Sarah Teichmann, a senior author from the Wellcome Sanger Institute and co-chair of the HCA Organising Committee, said: “As we’re building the Human Cell Atlas it is already being used to understand COVID-19 and identify which of our cells are critical for initial infection and transmission. This information can be used to better understand how coronavirus spreads. Knowing which exact cell types are important for virus transmission also provides a basis for developing potential treatments to reduce the spread of the virus.”
The global HCA Lung Biological Network continues to analyse the data in order to provide further insights into the cells and targets likely to be involved in COVID-19, and to relate them to patient characteristics.
Professor Sir Jeremy Farrar, Director of Wellcome, said: “By pinpointing the exact characteristics of every single cell type, the Human Cell Atlas is helping scientists to diagnose, monitor and treat diseases including COVID-19 in a completely new way. Researchers around the world are working at an unprecedented pace to deepen our understanding of COVID-19, and this new research is testament to this. Collaborating across borders and openly sharing research is crucial to developing effective diagnostics, treatments and vaccines quickly, ensuring no country is left behind.”
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Cell Analysis
Cell analyse advanced method allows us to explore and understand cells in their many facets. From single cell analysis to flow cytometry and imaging technology, cell analysis provides us with valuable insights into the structure, function and interaction of cells. Whether in medicine, biological research or pharmacology, cell analysis is revolutionizing our understanding of disease, development and treatment options.
Topic World Cell Analysis
Cell analyse advanced method allows us to explore and understand cells in their many facets. From single cell analysis to flow cytometry and imaging technology, cell analysis provides us with valuable insights into the structure, function and interaction of cells. Whether in medicine, biological research or pharmacology, cell analysis is revolutionizing our understanding of disease, development and treatment options.