Resistance can spread even without the use of antibiotics
Advertisement
antibiotic resistance does not spread only where and when antibiotics are used in large quantities, ETH researchers conclude from laboratory experiments. Reducing antibiotic use alone is therefore not sufficient to curtail resistance, and should be done in conjunction with measures to prevent infection with resistant germs.
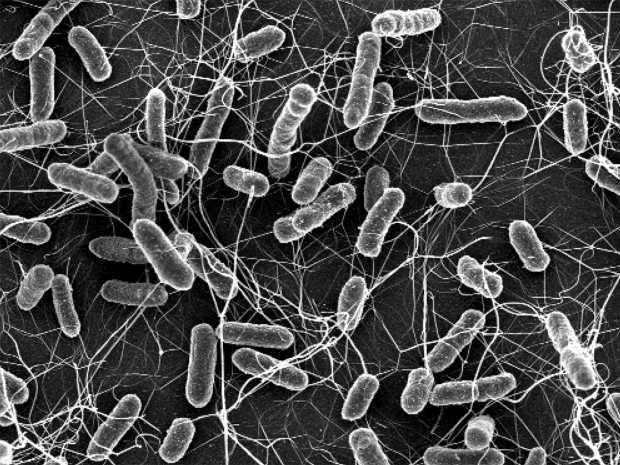
Salmonella causes diarrhoea in animals and humans. These bacteria become a particular public health concern if they are resistant to antibiotics (electron microscopic photograph).
Elektronenmikroskopisches Bild: ETH Zürich / Stefan Fattinger
Bacteria are becoming increasingly resistant to common antibiotics. Often, resistance is mediated by resistance genes, which can simply jump from one bacterial population to the next. It’s a common assumption that the resistance genes spread primarily when antibiotics are used, a rationale backed up by Darwin’s theory: only in cases where antibiotics are actually being used does a resistant bacterium have an advantage over other bacteria. In an antibiotic-free environment, resistant bacteria have no advantage. This explains why health experts are concerned about the excessive use of antibiotics and call for more restrictions on their use.
However, a team of researchers led by scientists from ETH Zurich and the University of Basel have now discovered an additional, previously unknown mechanism that spreads resistance in intestinal bacteria that is independent of the use of antibiotics. “Restricting the use of antibiotics is important and the indeed the right thing to do, but this measure alone is not sufficient to prevent the spread of resistance,” says Médéric Diard, who until recently held a post at ETH Zurich and is now a professor at the Biozentrum of the University of Basel. He continues, “If you want to control the spread of resistance genes, you have to start with the resistant microorganisms themselves and prevent these from spreading through, say, more effective hygiene measures or vaccinations.” Diard led the research project together with Wolf-Dietrich Hardt, Professor for Microbiology at ETH Zurich.
Two resistance mechanisms combined
Persistent bacteria, also known as persisters, are responsible for this newly discovered resistance-spreading mechanism. Scientists have known for some time that, just like bacteria that carry resistance genes, persisters can survive antibiotic treatment. They fall into a temporary, dormant state and can reduce their metabolism to a minimum, which prevents the antibiotics from killing them. In the case of salmonella, the bacteria become dormant when they penetrate the body tissue from inside the gut. Once they have invaded the tissue, the persisters can live there undetected for months before awakening from their dormant state. If the conditions are conducive to bacterial survival, the infection can flare up again.
Even if the persisters don’t cause a new infection, they can still have an adverse effect, as the scientists report in the journal Nature. In salmonella, a combination of the two resistance mechanisms is common: persisters that also carry small DNA molecules (plasmids) containing resistance genes.
Reservoir of genetic information
In experiments with mice, the researchers demonstrated that dormant salmonella in the gut can pass their resistance genes on to other individual bacteria of the same species and even to other species, such as E. coli from the normal intestinal flora. Their experiments showed that persisters are very efficient at sharing their resistance genes as soon as they awaken from their dormant state and encounter other bacteria that are susceptible to gene transfer. “By exploiting their persistent host bacterium, the resistance plasmids can survive for a prolonged period in one host before transferring into other bacteria. This speeds up their spread,” ETH professor Hardt explains. It’s important to note here that this transfer happens regardless of whether antibiotics are present or not.
The researchers now want to take their findings in mice and explore these more closely in livestock that frequently suffer from salmonella infections, such as pigs. The scientists also want to investigate whether it’s possible to control the spread of resistance in livestock populations with probiotics or with a vaccination against salmonella.