To use all functions of this page, please activate cookies in your browser.
my.bionity.com
With an accout for my.bionity.com you can always see everything at a glance – and you can configure your own website and individual newsletter.
- My watch list
- My saved searches
- My saved topics
- My newsletter
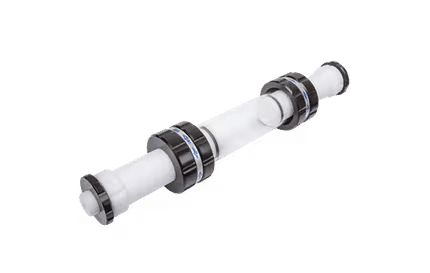
Site-directed mutagenesisSite-directed mutagenesis is a molecular biology technique in which a mutation is created at a defined site in a DNA molecule, usually a circular molecule known as a plasmid. In general, site-directed mutagenesis requires that the wild-type gene sequence be known. This technique is also known as site-specific mutagenesis or Oligonucleotide-directed mutagenesis. Product highlight
InventionAn important milestone in modern molecular biology came in 1978 with the first description of site-directed mutagenesis using oligonucleotides for in vitro synthesis of mutant DNA. So central has this technique become to all of biochemistry and molecular biology that Michael Smith, its pioneer, shared the Nobel Prize in Chemistry in October 1993 with Kary B. Mullis who developed PCR. In 1987 Kunkel et al. introduced an improvement to the technique that eliminated the need for phenotypic selection. The plasmid to be mutated would be transformed into an E. coli deficient in two genes, dUTPase & uracil deglycosidase. The former would prevent the breakdown of dUTP, a nucleotide that replaces dTTP in RNA, resulting in an abundance of the molecule, and deficiency in the latter would prevent the removal of dUTP from newly synthesized DNA. As the double-mutant E. coli replicates the up-taken plasmid, its enzymatic machinery incorporates the dUTP, resulting in a distinguishable copy. This copy is then extracted and incubated with an oligonucleotide containing the desired mutation, which attaches by base pair hydrogen bonding to the complementary wild-type gene sequence, as well as the Klenow enzyme, dNTPs, and DNA ligase. The reaction essentially replicates the dUTP-containing plasmid using as primer the oligonucleotide, giving a nearly identical copy. The essential differences being that the copy contains dTTP rather than dUTP, as well as the desired mutation. When the chimeric double-stranded plasmid, containing the dUTP, unmutated strand and the dTTP, mutated strand, is inserted into a normal, wild-type E. coli, the dUTP-containing strand is broken down, whereas the mutation-containing strand is replicated. Cassette MutagenesisThis type of mutagenesis involves the cleavage by a restriction enzyme (RE) at a site in the plasmid and subsequent ligation of an oligonucleotide containing the mutation in the gene of interest to the plasmid. Usually the RE that cuts at the plasmid and the oligonucleotide is the same, permitting sticky ends of the plasmid and insert to ligate to one another. Through PCRUsually focused on plasmid templates, the PCR method involves the use of oligonucleotide "primers" containing the desired mutation. The theory is woven into the PCR method: As the primers are the ends of newly-synthesized strands, should there be a mis-match during the first cycle in binding the template DNA strand, after that first round, the primer-based strand (containing the mutation) would be at equal concentration to the original template. After successive cycles, it would exponentially grow, and after 25, would outnumber the original, unmutated strand in the region of 8 million : 1, resulting in a nearly homogeneous solution of mutated amplified fragments. Despite the fact that the PCR product is overwhelmingly composed of mutated plasmid, the template DNA is typically eliminated by enzymatic digestion with Dpn1, a restriction enzyme which cleaves only Dam methylated DNA. The template, which is derived from an alkaline lysis plasmid preparation and therefore is methylated, is destroyed in this step, but the mutated plasmid is preserved because it was generated in vitro and is unmethylated as a result. See also
Categories: Mutation | Molecular genetics |
|
| This article is licensed under the GNU Free Documentation License. It uses material from the Wikipedia article "Site-directed_mutagenesis". A list of authors is available in Wikipedia. |