Berthold Technologies
Improved experimental setup for analysis of circadian rhythms using the nightshade
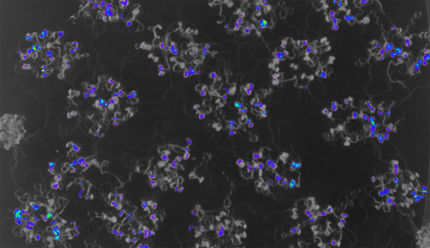
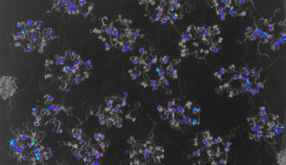
An improved Plant In Vivo Imaging protocol for circadian rhythms experiments using the NightShade
Endogenous biological clocks drive daily rhythms enabling organisms to anticipate environmental changes as well as to coordinate and adapt their physiology in a synchronized manner. Research on circadian rhythms benefits from real-time monitoring of reporter lines in which the promoter of a gene of interest drives the expression of luciferase pGENE::LUC+ in combination with sensitive imaging systems [1]. However, in multicellular organisms, circadian clocks are naturally variable at individual, tissue as well as cellular level [2, 3], culminating in noisy or inaccurate data. Therefore, robustness is required to accurately address key questions in circadian biology. For this purpose, we developed a simple protocol for circadian rhythms experiments with Arabidopsis thaliana reporter lines using the NightShade LB 985. Our experimental setup improves data quality, reduces luminescence variation between replicates and highly correlates with modelling predictions.