Team develops strategy to determine how non-coding variants contribute to disease risk
Researchers describes a strategy for meeting one of today's most significant challenges in genomic medicine - determining whether a specific DNA variant in the non-protein-coding genome is the actual disease-causing variant of an associated disease risk. The report from a multi-institutional team study led by investigators at the Dana-Farber Cancer Institute (DFCI), Massachusetts General Hospital (MGH), and the Keck School of Medicine at the University of Southern California describes a procedure called CAUSEL (Characterization of Alleles Using Editing of Loci) that uses epigenome- and genome-editing tools to determine functional causality of disease-associated variants in the non-coding genome and to study the mechanisms by which those variants contribute to disease.
"This is a good example of how the intersection of cutting-edge genetic and epigenetic profiling with the latest genome- and epigenome-editing technologies can be used to advance our understanding of how sequence variants can impact diseases," says J. Keith Joung, MD, PhD, associate chief for Research in the MGH Department of Pathology and a co-senior author of the report. "We believe that this is an important next frontier for advancing diagnosis and treatment of diseases influenced by genetics and that CAUSEL provides a blueprint for how to proceed with these types of studies."
Co-senior author Matthew Freedman, MD, of the DFCI Center for Cancer Genome Discovery and Center Functional Cancer Epigenetics explains that the effort to determine the precise DNA variants that increase disease risk and how they do so is extremely complicated. While the genetic mapping approach known as linkage analysis has enabled the identification of DNA variants in protein-coding genes - like the BRCA1 and 2 genes that, when mutated, cause a clearly increased, inherited risk of breast and ovarian cancer - those variants account for approximately 5 percent of cases of inherited disease risk. The other 95 percent appears to be predominantly influenced by variants mapping to non-protein-coding regulatory elements that control the levels at which protein-coding genes are expressed.
The CAUSEL procedure or 'pipeline' consists of five steps:
- genetic fine mapping to identify candidate variants,
- epigenomic profiling, in which the candidate variants are intersected with epigenetic data to identify which are most likely to cause the condition,
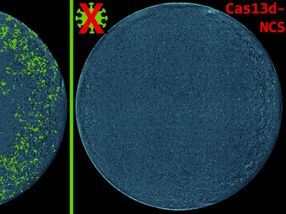
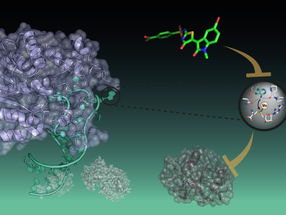
- epigenomic editing, in this case using reagents developed in Joung's laboratory, to confirm whether or not the candidate variants may possess regulatory capacity,
- genome editing, to create cell lines with all possible genotypes of the candidate variants,
- phenotypic analysis of those cell lines, to evaluate functional differences relevant to the disease or condition of interest.
The researchers tested this pipeline on a region of chromosome 6 that previous studies have associated with increased prostate cancer risk, probably by controlling expression levels of RFX6, a regulatory protein involved with tumor-associated properties. Fine-mapping that region identified 27 candidate variants, all associated with increased prostate cancer risk, and epigenomic profiling identified one as most likely to be relevant. Epigenomic editing performed at that site, called rs339331, confirmed that the region it lies in could potentially regulate RFX6 expression, so the investigators created three cell lines - one with two copies of the disease-associated variant or allele, one with two copies of the 'normal' version, and one with a copy of both versions.
Analysis of these cell lines revealed that, while cells with two normal alleles had the appearance of normal cells, both lines containing the cancer-associated variant had an appearance more typical of cancer cells. Cells carrying two cancer-related alleles were more likely to adhere to surfaces, a property typical of cancer cells, and those cells lines also exhibited changes in the expression of genes involved with androgen signaling, a pathway known to be critical in the risk for and treatment of prostate cancer.